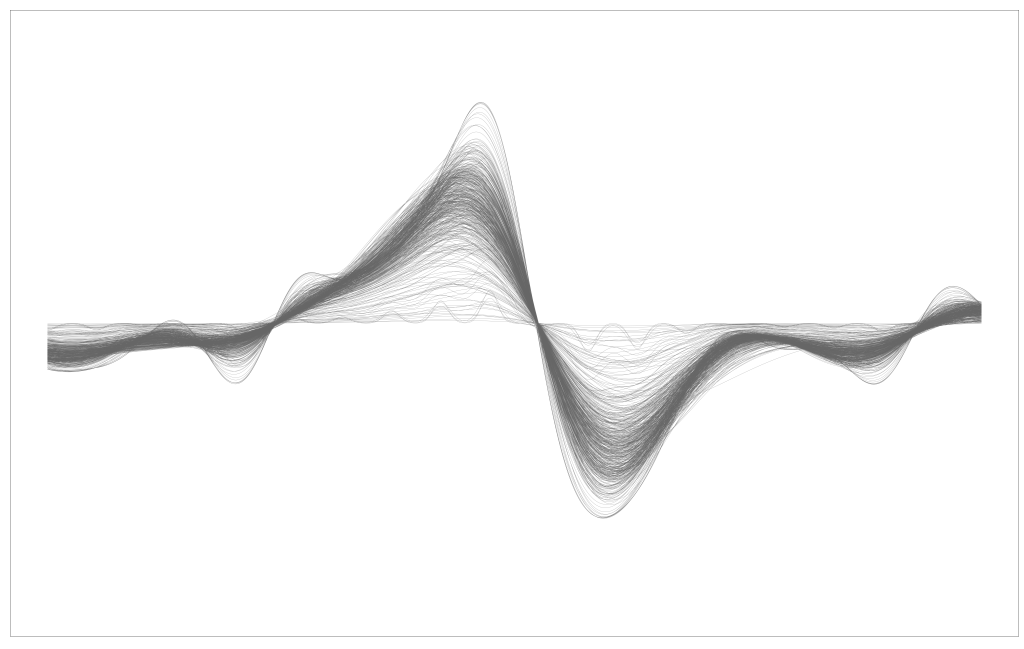
Prior to starting at SAS, I was still consulting and working for Michigan. I wanted to make a logo for Likely LLC, to make it appear more credible. Models are meaningless without good advertising and visualizations, to non-statisticians. I was attempting to make an L with mathematics and machine learning/statistics. So I simulated a Paul Wavelet, added some additive white Gaussian noise (AWGN), and then fit a Gaussian process regression model and plotted a few realizations. It didn’t quite look like an L, but I had to prepare for an interview. Admittedly, probably, the most useless application of these tools is to make art. Below is some of the code. I had to use a few different programming languages. Python, Stan, R. Python I can’t find. You can see me experimenting with different colors, but you can also play around with the amount of realizations and stuff.
setwd("/home/andre/likely/logo/")
library(rstan);
set.seed(1);
x = read.csv("paul2_wavelet.csv");
x = as.matrix(x);
x = x[seq(1,length(x), by = 30)]; ## evenly spaced
N = length(x);
## sample by index
# N2 = 20;
# x_sub = sort(sample(1:N, size = N2));
# plot(1:N, x);
# plot(1:N2, x[x_sub]);
x_eps = x + rnorm(N, 0, .1)
plot(1:N, x_eps)
N = N;
D = 1;
y = as.vector(x_eps);
x = as.matrix(1:length(y));
N_pred = 1000;
x_pred = as.matrix(seq(min(x) - 10, max(x) + 10, length.out = 1000));
rstan::stan_rdump(list = c("N", "D", "x", "y", "N_pred", "x_pred"), file = "paul_sim.staninput");
####
csv = read_stan_csv("output.csv");
samples = rstan::extract(csv);
for (i in 101:200){
plot(samples$f_pred[i, 0:1000],ylim = c(-1,1), type = "l", yaxt = "n", xaxt = "n", xlab = "", ylab = "", col = "grey")
par(new = T)
}
for (i in 1:300){
plot(samples$f_pred[i, 0:1000],ylim = c(-1,1), type = "l", yaxt = "n", xaxt = "n", xlab = "", ylab = "", col = "grey")
par(new = T)
}
for (i in 1:500){
plot(samples$f_pred[i, 0:1000],ylim = c(-1,1), type = "l", yaxt = "n", xaxt = "n", xlab = "", ylab = "", col = "grey")
par(new = T)
}
for (i in 1:500){
plot(samples$f_pred[i, 0:1000],ylim = c(-1,1), type = "l", yaxt = "n", xaxt = "n", xlab = "", ylab = "", col = "gray40")
par(new = T)
}
for (i in 1:500){
plot(samples$f_pred[i, 0:1000],ylim = c(-1,1), type = "l", yaxt = "n", xaxt = "n", xlab = "", ylab = "", col = "gray50")
par(new = T)
}functions {
vector gp_pred_rng(vector[] x_pred,
vector y1, vector[] x,
real magnitude, real length_scale,
real sigma) {
int N = rows(y1);
int N_pred = size(x_pred);
vector[N_pred] f2;
{
matrix[N, N] K = add_diag(gp_exp_quad_cov(x, magnitude, length_scale),
sigma);
matrix[N, N] L_K = cholesky_decompose(K);
vector[N] L_K_div_y1 = mdivide_left_tri_low(L_K, y1);
vector[N] K_div_y1 = mdivide_right_tri_low(L_K_div_y1', L_K)';
matrix[N, N_pred] k_x_x_pred = gp_exp_quad_cov(x, x_pred, magnitude, length_scale);
f2 = (k_x_x_pred' * K_div_y1);
}
return f2;
}
}
data {
int<lower=1> N;
int<lower=1> D;
vector[D] x[N];
vector[N] y;
int<lower=1> N_pred;
vector[D] x_pred[N_pred];
//real x_pred[N_pred];
}
transformed data {
vector[N] mu;
mu = rep_vector(0, N);
}
parameters {
real<lower=0> magnitude;
real<lower=0> length_scale;
real<lower=0> sig;
real<lower=0> sigma;
}
transformed parameters {
matrix[N, N] L_K;
{
matrix[N, N] K = gp_exp_quad_cov(x, magnitude, length_scale);
K = add_diag(K, square(sigma));
L_K = cholesky_decompose(K);
}
}
model {
magnitude ~ normal(0, 1);
length_scale ~ normal(0, 1);
sig ~ normal(0, 1);
sigma ~ normal(0, 1);
y ~ multi_normal_cholesky(mu, L_K);
}
generated quantities {
vector[N_pred] f_pred = gp_pred_rng(x_pred, y, x, magnitude, length_scale, sigma);
vector[N_pred] y_pred;
for (n in 1:N_pred) y_pred[n] = normal_rng(f_pred[n], sigma);
}
Leave a comment